IDmyBee online identification tools rely on the Xper3 platform (website).
Xper3 is a free, online platform to manage biodiversity data, including databases for identification tools. It enables the collaborative development of identification tools.
Using those online identification tools is easy and intuitive. Here, we will see how to use this platform for insect identification using the IDmyBee-Genera tool on computer as an example.
Xper3 is a free, online platform to manage biodiversity data, including databases for identification tools. It enables the collaborative development of identification tools.
Using those online identification tools is easy and intuitive. Here, we will see how to use this platform for insect identification using the IDmyBee-Genera tool on computer as an example.
Overview of the ID tool.
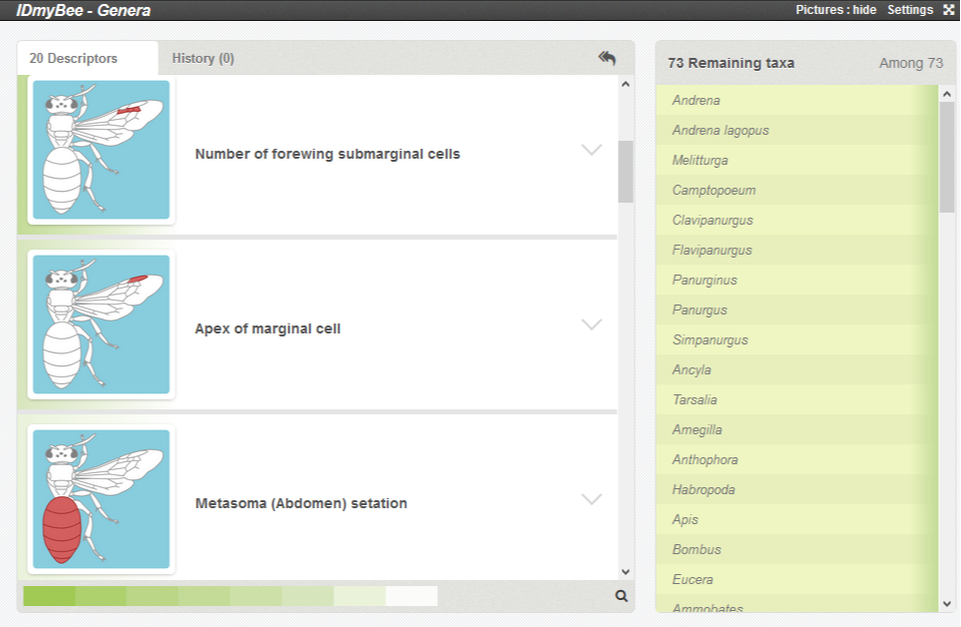
Xper3 offers different options, but the standard view is the following:
On the left, the descriptors : characters that you have to look for on your specimen.
On the right, a panel with the list of taxa included in the tool.
On the right, a panel with the list of taxa included in the tool.
Select a descriptor to start identifying your insect.
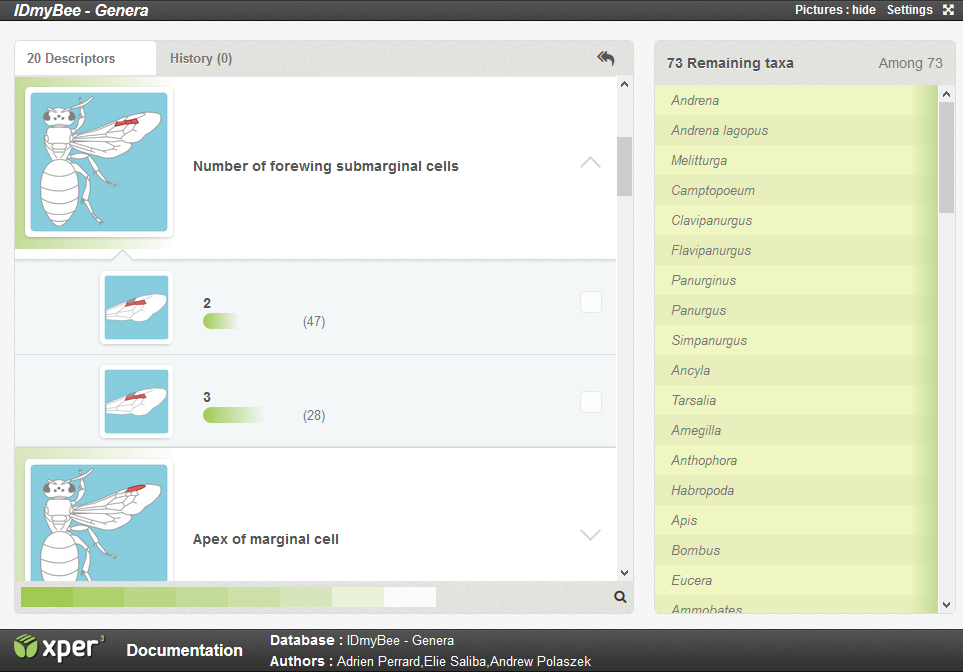
You can select any descriptor on the list. They are presented in a semi-organized order: the more discriminant ones are at the beginning of the list. You may want to start by one of those, but you don't have to.
When you click on a descriptor, the different possible states of the character will appear. In the example above, the descriptor 'Number of forewing submarginal cells' has 2 possible states : 2 (submarginal cells) or 3.
Next to each state, you have the number of taxa with the corresponding state.
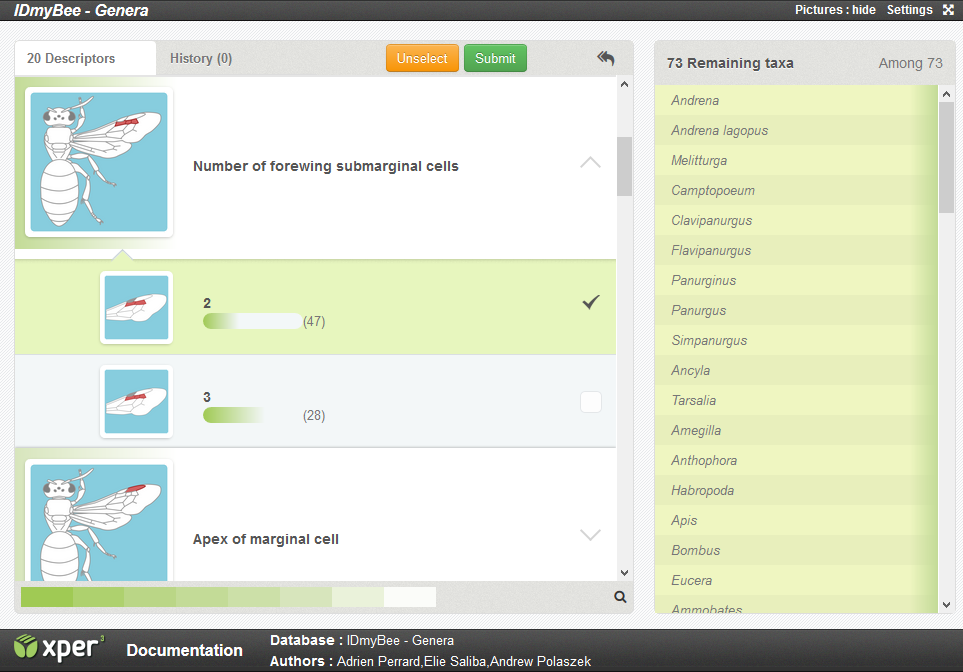
Make your choice by selecting the state corresponding to what you can see on your specimen.
When you select a state, two buttons (Unselect & Submit) appear at the top of the screen.
To validate your choice, you need to click on 'Submit'.
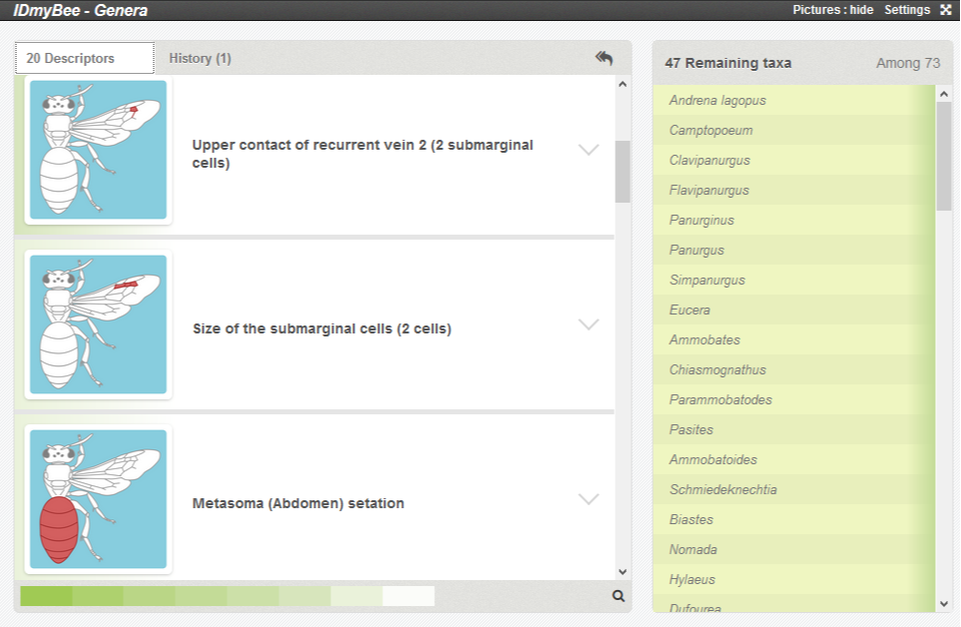
This action will remove the taxa which do not present this state from the list and let you choose another descriptor to use.
Below, you can see in the taxa panel that only 47 of the 73 taxa remain when 2 submarginal cells are validated.
Next to each state, you have the number of taxa with the corresponding state.
Make your choice by selecting the state corresponding to what you can see on your specimen.
When you select a state, two buttons (Unselect & Submit) appear at the top of the screen.
To validate your choice, you need to click on 'Submit'.
This action will remove the taxa which do not present this state from the list and let you choose another descriptor to use.
Below, you can see in the taxa panel that only 47 of the 73 taxa remain when 2 submarginal cells are validated.
IMPORTANT: Xper3 works by eliminating taxa with contradicting descriptions. It does not mean that the remaining taxa have the described state : some may remain because they have no state described.
It can happen if their description is incomplete or because the descriptor cannot be applied to them.
It can happen if their description is incomplete or because the descriptor cannot be applied to them.
You can doubt and still make progress in the identification
Xper3 let you select more than one descriptor state. So if you are not sure of your observation, you don't have to choose between 2 states and select both of them. It can still be useful if the descriptor has more than 2 states.
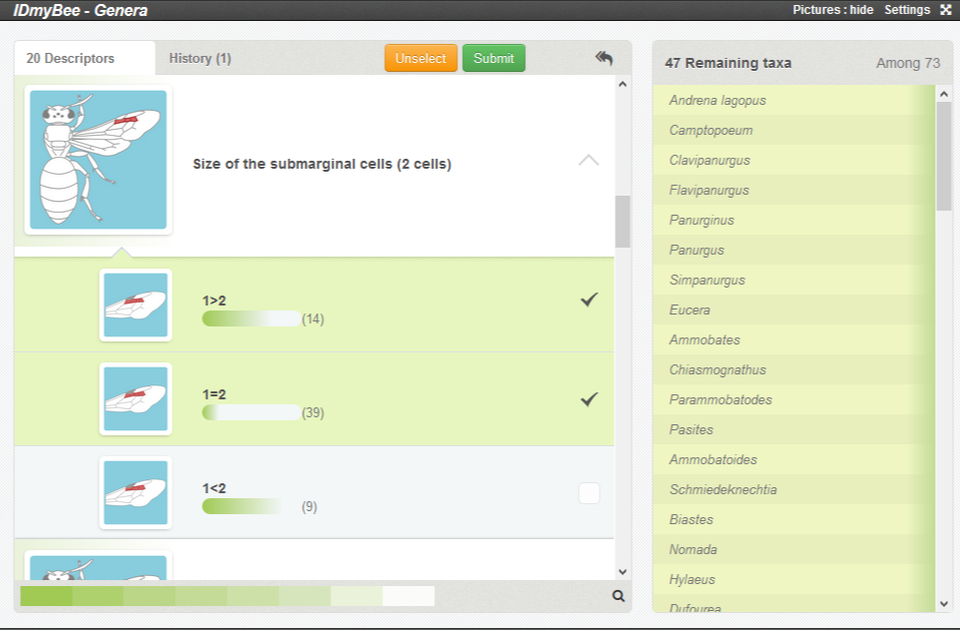
In the example below, the relative size of the two submarginal cells may be difficult to assess, so both 'cell 1 larger than cell 2' and 'cell 1 and 2 of equivalent sizes' are selected.
Such an option will remove only taxa with cell 2 distinctly larger than cell 1 from the list.
In the example below, the relative size of the two submarginal cells may be difficult to assess, so both 'cell 1 larger than cell 2' and 'cell 1 and 2 of equivalent sizes' are selected.
Such an option will remove only taxa with cell 2 distinctly larger than cell 1 from the list.
The eliminated taxa do not disappear
When you submit different observations, the list of remaining taxa will reduce, but you can still see which taxa corresponds more or less to the description of your specimen.
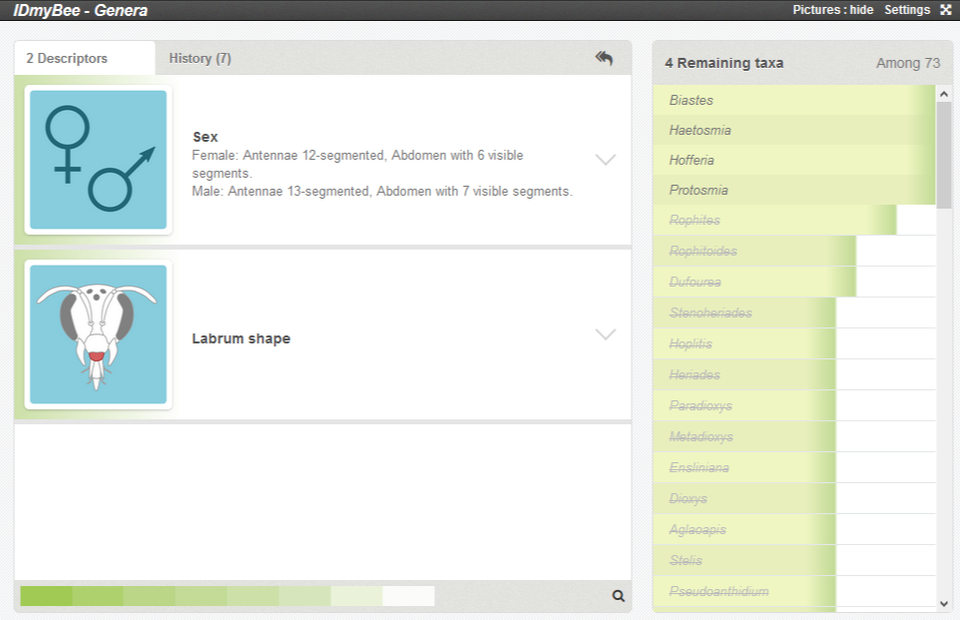
In the example below, after more descriptions submitted, only 4 taxa remain entirely congruent with the description of the specimen. You can see than the list of the eliminated taxa is not random: Rophites appears first, with a larger green band than the other eliminated taxa.
The colour band in the taxa list represent the percentage of congruence between the description of the specimen and the taxon. This band is full (100%) for the remaining taxa, and decreases for eliminated taxa. This feature may be interesting if you notice some discrepancies between your identification result and your specimen.
In the example below, after more descriptions submitted, only 4 taxa remain entirely congruent with the description of the specimen. You can see than the list of the eliminated taxa is not random: Rophites appears first, with a larger green band than the other eliminated taxa.
The colour band in the taxa list represent the percentage of congruence between the description of the specimen and the taxon. This band is full (100%) for the remaining taxa, and decreases for eliminated taxa. This feature may be interesting if you notice some discrepancies between your identification result and your specimen.
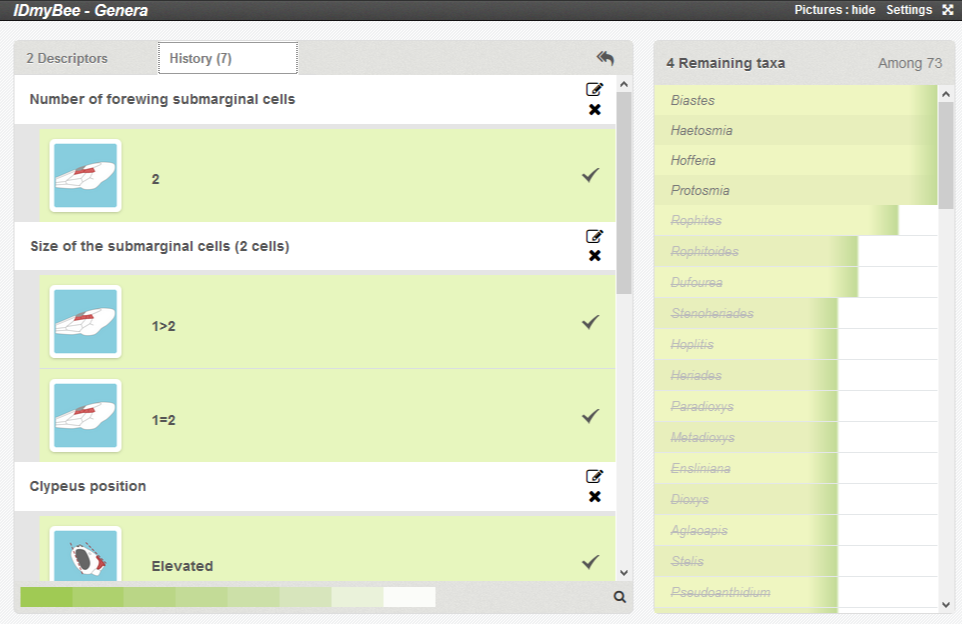
You can even go back to the history of your identification process by clicking on the 'History' panel on the upper left, above the descriptor list.
This panel let you review the states you selected, and modify your description if required.
This panel let you review the states you selected, and modify your description if required.
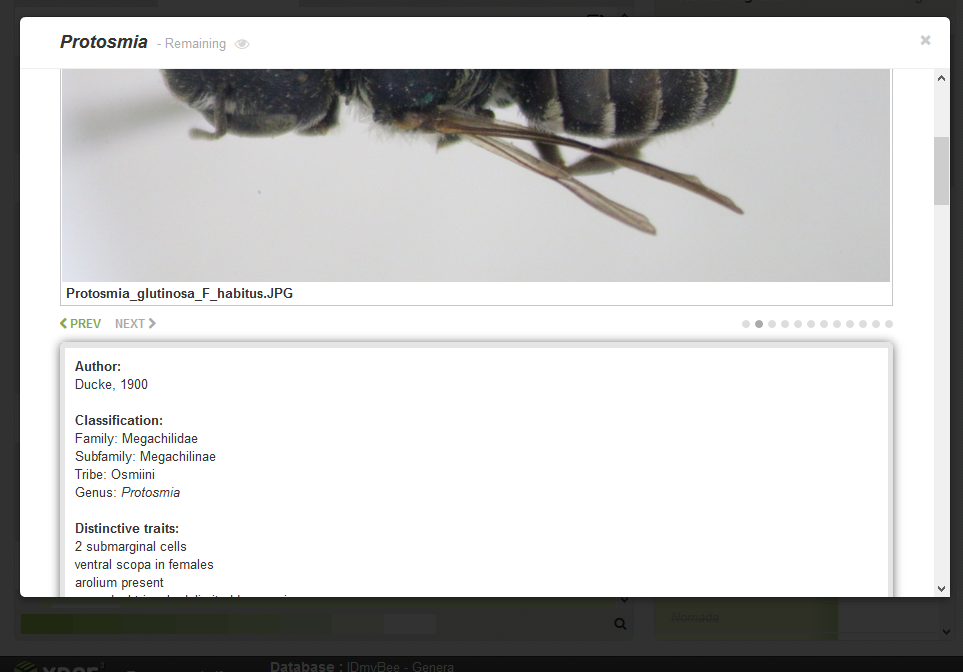
Finally, you can click on the name of a taxon in the right panel (remaining or eliminated) to open a page with information about the taxon.
There you can find pictures (you can navigate between pictures with the buttons 'prev' and 'next' at the bottom left), information about the taxon, a link to the IDmyBee page and the list of descriptors with their states coded for this taxon.
There you can find pictures (you can navigate between pictures with the buttons 'prev' and 'next' at the bottom left), information about the taxon, a link to the IDmyBee page and the list of descriptors with their states coded for this taxon.
That's it! Now you should be able to identify bees using our Xper3 tools!
Please contact us if you encounter trouble or if you need more information.